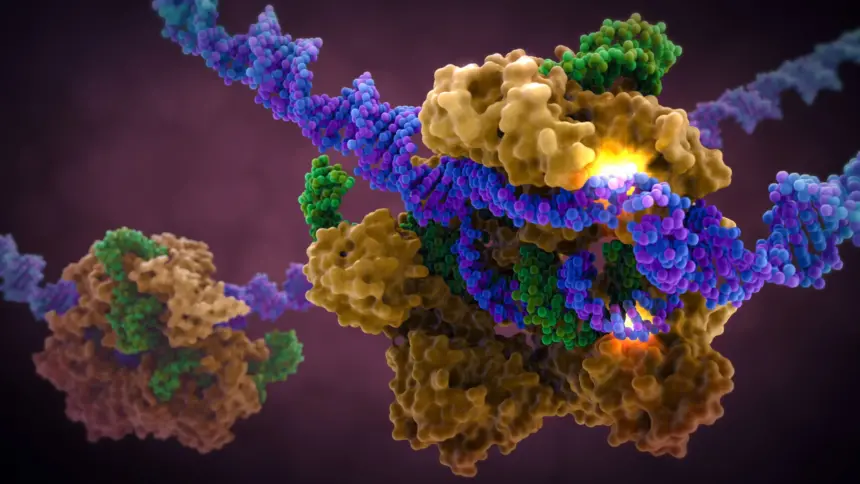
CRISPR, an acronym for “Clustered Regularly Interspaced Short Palindromic Repeats”, is a name given to certain sequences of DNA found in prokaryotes, organisms with no nucleus. These sequences are actually part of the genome of a virus which had infected the bacterium before. The bacterium keeps the viral DNA segment, called a CRISPR sequence, and combines it with an enzyme called Cas9 to form the CRISPR-Cas9 complex, which then detects and deactivates the virus genome if it is ever present again. This complex is the basis of modern gene editing technology and has been used to treat diseases, develop biological products such as insulin and is a significant focus in biology.
To understand how CRISPR-Cas9 gene editing works, we have to understand the two main components of the technology: The CRISPR sequences and the Cas9 protein.
When a virus invades a bacterium, in most cases, the virus multiplies and kills its host. However, in some cases, the bacterium lives and integrates the virus’ DNA into its genome. This is what makes up the CRISPR sequences, a mix of “spacers”, viral DNA, and “repeats”, short DNA segments that separate the parts of viral DNA.
There is another type of important gene in this scenario, the “Cas (CRISPR-associated)” genes, that encode special “Cas” enzymes. If the CRISPR sequences are guidelines, then the Cas proteins are the scissors of the bacterium. They serve to detect the genes encoded in the “spacers” and cuts those parts if they are found in the bacterium again.
Despite the knowledge we have of this system, it’s relatively new. It was first discovered in 1987, albeit accidentally, when researchers cloned a part of a CRISPR sequence. It was realized near the end of the 1990s that this was a common case, so common in fact that scientists decided to give it a name, and in 2002, the term “CRISPR” was used for the first time. More breakthroughs were made in the 2000s when similarities between the CRISPR sequences and virus genes were discovered and it was suggested that it was used by bacteria to defend against viruses. A 2007 experiment found that when a colony of yeast was infected with viruses, all but a certain amount of cells died. However, the remaining cells proved to be resistant to the viruses and produced resistant offspring as well. This resistance disappeared when the previously discovered “CRISPR sequences” were removed from the cell, proving CRISPR’s use as a sort of acquired immune system. Not long after these breakthroughs, scientists began to experiment and in 2012, published papers theorizing that CRISPR could be used for gene editing. This proved to be revolutionary, and soon after multiple papers by many different research groups were published on the topic, depicting how to use the system to edit genes, starting the age of CRISPR.

Today, the technology is still being improved, however in recent years there have been ethical debates about the usage & safety of CRISPR. Some institutions have put restrictions in place regarding gene modification and specifically CRISPR systems, while others have remained neutral about such applications.
Despite concerns, it is undeniable that CRISPR systems have a lot of potential. In the future, we may see them become a staple of agriculture, medicine, biotechnology and many more fundamental areas.